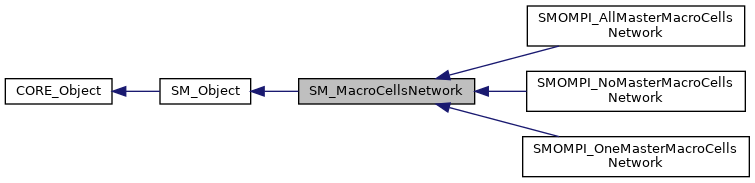
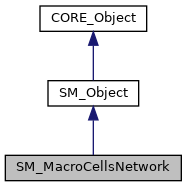
This class is describes a macro cell network. More...
#include <SM_MacroCellsNetwork.h>
Public Member Functions | |
| virtual tMemSize | getMemorySize () const |
| return the memory size of the class and the memory size of all its attributes/associations More... | |
| virtual tMemSize | getContentsMemorySize () const |
| return the memory size of the included associations More... | |
| virtual CORE_UniquePointer< SM_MacroCellsNetwork > | newInstance () const =0 |
| create a New instance of this More... | |
| void | setMacroCellMargin (const tReal &w) |
| set margin More... | |
| const tReal & | getMacroCellMargin () const |
| get margin More... | |
| void | setMacroCellSize (const std::array< tReal, SM_Constants::DIM > &H) |
| set the macro cell size in angstrom More... | |
| void | setMacroCellSize (const std::vector< tReal > &H) |
| set the macro cell size in angstrom More... | |
| const std::array< tReal, SM_Constants::DIM > & | getMacroCellSize () const |
| get the macro cell size | |
| void | setMacroCellsNumber (const tInteger &n) |
| set the number of macro cells | |
| const tInteger & | getMacroCellsNumber () const |
| get the number of macro cells | |
| const SM_RealField & | getMacroCellsMassCenter () const |
| get the mass center of the macro cells for reading | |
| SM_RealField & | getMacroCellsMassCenter () |
| get the mass center of the macro cells for writing | |
| const std::valarray< tInteger > & | getMacroCellsList () const |
| get the index of macro cell containing each particle p | |
| std::valarray< tInteger > & | getMacroCellsList () |
| get the index of macro cell containing each particle p | |
| const std::valarray< tReal > & | getMacroCellsVolume () const |
| get volume of macro cells in Angstrom | |
| std::valarray< tReal > & | getMacroCellsVolume () |
| get volume of macro cells in Angstrom | |
| const tReal & | getMacroCellVolume (const tInteger &m) const |
| get volume of macro cell at index m in Angstrom More... | |
| const tInteger & | getMacroCellIndex (const tInteger &p) const |
| get the local index of the macro cell containing particle p More... | |
| virtual void | computeMacroCells (const SM_Material &material, const SM_Network &network) |
| compute the macro cells of the network More... | |
| virtual void | computeMacroCellsMassCenter (const SM_Material &material, const SM_Network &network) |
| compute the macro cells of the network More... | |
| virtual tString | toString () const override |
| return string representaton of the operator | |
 Public Member Functions inherited from SM_Object Public Member Functions inherited from SM_Object | |
| SM_Object (void) | |
| create | |
| virtual | ~SM_Object (void) |
| destroy | |
 Public Member Functions inherited from CORE_Object Public Member Functions inherited from CORE_Object | |
| template<class T > | |
| std::shared_ptr< T > | getSharedPointer () |
| return the shared pointer for this More... | |
| template<class T > | |
| std::shared_ptr< const T > | getConstSharedPointer () const |
| return a const shared pointer for this More... | |
| template<class T > | |
| tBoolean | isInstanceOf () const |
| test if the clas T is an instance of this class More... | |
| tString | getClassName () const |
| return the name of the class More... | |
| tString | getPointerString () const |
| retrun the pointer of the class as a string More... | |
| tString | getIdentityString () const |
| retrun the string identification of the class More... | |
Protected Member Functions | |
| SM_MacroCellsNetwork (void) | |
| create a network class | |
| virtual | ~SM_MacroCellsNetwork (void) |
| destroy | |
| virtual void | computeBoundingBoxMacroCells (const SM_Material &material, const SM_Network &network, const std::array< tReal, SM_Constants::DIM > &P, const std::array< tReal, SM_Constants::DIM > &Q) |
| compute the macro cells of the network More... | |
 Protected Member Functions inherited from CORE_Object Protected Member Functions inherited from CORE_Object | |
| CORE_Object () | |
| build an instance of the object | |
| virtual | ~CORE_Object () |
| destroy the instance of object std | |
Static Protected Member Functions | |
| static tInteger | ComputeMacroCells (const SM_Material &material, const SM_Network &network, const std::array< tReal, SM_Constants::DIM > &P, const std::array< tReal, SM_Constants::DIM > &Q, const std::array< tReal, SM_Constants::DIM > ¯oCellSize, const tReal &margin, const tReal &epsVW, std::valarray< tReal > &MCsVolume, std::valarray< tInteger > ¯oCellsList) |
| compute the macro cells points and magnetization More... | |
| static void | EliminateEmptyMacroCells (const tReal &epsVM, std::valarray< tReal > &gMCVolume, std::valarray< tReal > &lMCVolume, std::valarray< tInteger > &gMCLocalIndices) |
| eliminate empty macro cells form lists More... | |
| static void | BuildParticlesListOrderedPerMacroCells (const SM_Network &network, const tInteger &nMacroCells, const std::valarray< tInteger > ¯oCellsPList, std::valarray< tInteger > &particlesMCList, std::valarray< tInteger > &particlesMCListOffset) |
| build the particles list for all not empty macro cells More... | |
| static void | ConvertIndices (const std::valarray< tInteger > &indicesConverter, std::valarray< tInteger > &indicesList) |
| convert indices More... | |
| static void | ConvertIndices (const std::valarray< tInteger > &indicesConverter, const tInteger &n, tInteger *iIndicesList) |
| convert indices More... | |
| static void | ComputeMacroCellX (const tInteger &nParticles, const tReal *iX, const tInteger *iMacroCellsList, SM_RealField &C, std::valarray< tInteger > &nMCParticles) |
| compute the sum of coordinantes of the particles in each macro cell More... | |
| static void | MeanX (const tInteger &n, const tInteger *iNs, tReal *iX) |
| compute the macro cells points and magnetization More... | |
Additional Inherited Members | |
 Static Public Member Functions inherited from CORE_Object Static Public Member Functions inherited from CORE_Object | |
| static tBoolean | EnableMemoryStack (const tBoolean &isMemoryChecked) |
| enable the memory stack More... | |
| static void | EnableMemoryStack () |
| enable the memory stack | |
| static void | DisableMemoryStack () |
| disable the memory stack | |
| static tBoolean | IsMemoryStackEnabled () |
| return trur if the memory stack is enabled | |
| static tString | MemoryStackToString () |
| get the memory stack in string More... | |
| static tIndex | GetRegisteredClassesNumber () |
| get the memory stack in string More... | |
Detailed Description
This class is describes a macro cell network.
- Version
- 2.0
Member Function Documentation
◆ BuildParticlesListOrderedPerMacroCells()
|
staticprotected |
build the particles list for all not empty macro cells
- Parameters
-
[in] network network pof particles [in] nMacroCells number of not empty macro cells [in] macroCellsPList index of macro cell for each particle [out] particlesMCList particles in a list ordred per macro cells [out] particlesMCListOffset : index of the first particle of each macro cell within the particlesMCList list
◆ computeBoundingBoxMacroCells()
|
protectedvirtual |
compute the macro cells of the network
- Parameters
-
[in] material :material data [in] network : network of the macro cells [in] P min point of the box to compute macro cells [in] Q max point of the box to compute macro cells
It computes the none empty macro cells list
Reimplemented in SMOMPI_OneMasterMacroCellsNetwork, and SMOMPI_AllMasterMacroCellsNetwork.
◆ computeMacroCells()
|
inlinevirtual |
compute the macro cells of the network
- Parameters
-
[in] material :material data [in] network : network of the macro cells
Reimplemented in SMOMPI_OneMasterMacroCellsNetwork, SMOMPI_NoMasterMacroCellsNetwork, and SMOMPI_AllMasterMacroCellsNetwork.
◆ ComputeMacroCells()
|
staticprotected |
compute the macro cells points and magnetization
- Parameters
-
[in] material :material data [in] network : network of the particle [in] P min point of the bouding box [in] Q max point of the bouding box [in] macroCellSize size of the macro cells in Angstrom [in] margin margin to shift the macro cells grid [in] epsVW : minimum volume under which the macro cell is supposed to be empty [out] MCsVolume : volume of each macro cells expressed in Angstrom whith global indices \( MCsVolume[mc]=V_a.card{p | p \in mc}, V_a \) is the atomic volume [out] macroCellsList list of macro cells for each particles
- Returns
- the number of not empty macro cells
◆ computeMacroCellsMassCenter()
|
virtual |
compute the macro cells of the network
- Parameters
-
[in] material :material data [in] network : network of the macro cells
Reimplemented in SMOMPI_OneMasterMacroCellsNetwork, and SMOMPI_AllMasterMacroCellsNetwork.
◆ ComputeMacroCellX()
|
staticprotected |
compute the sum of coordinantes of the particles in each macro cell
- Parameters
-
[in] nParticles number of particles [in] iX values of coordinates of the nParticles [in] iMacroCellsList : index of macro cell per each particle [out] C : sum of the coodrinate of p within each macro cell [out] nMCParticles number of particles in each macro cell
◆ ConvertIndices() [1/2]
|
inlinestaticprotected |
convert indices
- Parameters
-
[in] indicesConverter : converter map from set E into set F [in] n : size of the list [in,out] iIndicesList : begin iterator on list of indices into E in input, list of indices into F in input
◆ ConvertIndices() [2/2]
|
inlinestaticprotected |
convert indices
- Parameters
-
[in] indicesConverter : converter map from set E into set F [in,out] indicesList : list of indices into E in input, list of indices into F in input
◆ EliminateEmptyMacroCells()
|
staticprotected |
eliminate empty macro cells form lists
- Parameters
-
[in] epsVM : minimum volume under which the macro cell is supposed to be empty [in,out] gMCVolume : volume of each macro cell with global indice [out] lMCVolume : volume of each macro cell with local indice within not empty macro cells [out] gMCLocalIndices map betwrn global indices of macro cell in the bouinding box grid to local index of macro cell within not empty macro cells list
◆ getContentsMemorySize()
|
inlinevirtual |
return the memory size of the included associations
- Returns
- the memory size of the storage in bytes 1 Kb = 1024 bytes 1 Mb = 1024 Kb 1 Gb = 1024 Mb 1 Tb = 1024 Gb 1 Hb = 1024 Tb
Reimplemented from CORE_Object.
Reimplemented in SMOMPI_OneMasterMacroCellsNetwork, SMOMPI_NoMasterMacroCellsNetwork, and SMOMPI_AllMasterMacroCellsNetwork.
◆ getMacroCellIndex()
|
inline |
get the local index of the macro cell containing particle p
- Parameters
-
[in] p : index of the particle
- Returns
- the index of macro cell
◆ getMacroCellMargin()
|
inline |
get margin
- Returns
- margin to increase the bounding box to be sure the bounding box of the domain is inside the bouding box of the macro cells
◆ getMacroCellVolume()
|
inline |
get volume of macro cell at index m in Angstrom
- Parameters
-
[in] m : idex of the macro cell
◆ getMemorySize()
|
inlinevirtual |
return the memory size of the class and the memory size of all its attributes/associations
- Returns
- the memory size of the class and the memory size of its attributes/associations in bytes The mamory size is :
- the added size of the base classes which contains:
- the primary attributes size depends on the order: (first delare the smallest attributes size
- all virtual functions costs <pointer-size> (4 32xor 8 64x) bytes by virtual function
- virtual inherihtance will increase of (4 or 8) bytes
- we add the size of the contains values of the attributes : for example the size of a string is the length of the string 1 octet = 1 byte 1 Ko = 1024 bytes 1 Mo = 1024 Ko 1 Go = 1024 Mo
- the added size of the base classes which contains:
Reimplemented from SM_Object.
Reimplemented in SMOMPI_OneMasterMacroCellsNetwork, SMOMPI_NoMasterMacroCellsNetwork, and SMOMPI_AllMasterMacroCellsNetwork.
◆ MeanX()
|
inlinestaticprotected |
compute the macro cells points and magnetization
- Parameters
-
[in] n number of elements [in] iNs number of samples per element [in,out] iX IN: sum of samples values OUT : mean of samples values
◆ newInstance()
|
pure virtual |
create a New instance of this
- Returns
- an unique pointer to the instance
Implemented in SMOMPI_OneMasterMacroCellsNetwork, SMOMPI_NoMasterMacroCellsNetwork, and SMOMPI_AllMasterMacroCellsNetwork.
◆ setMacroCellMargin()
|
inline |
set margin
- Parameters
-
[in] w : margin to increase the bounding box to be sure the bounding box of the domain is inside the bouding box of the macro cells
◆ setMacroCellSize() [1/2]
|
inline |
set the macro cell size in angstrom
- Parameters
-
[in] H : size of the macro cell in angstrom
◆ setMacroCellSize() [2/2]
|
inline |
set the macro cell size in angstrom
- Parameters
-
[in] H : size of the macro cell in angstrom
The documentation for this class was generated from the following files:
- /home/despreau/Developpement/CPP/devcpp20/include/stochmagnet/operators/fields/SM_MacroCellsNetwork.h
- /home/despreau/Developpement/CPP/devcpp20/src/stochmagnet/operators/fields/SM_MacroCellsNetwork.cpp